Show Summary Copy Number Profile for Sample Groups
Source:R/show_cn_group_profile.R
show_cn_group_profile.RdShow Summary Copy Number Profile for Sample Groups
show_cn_group_profile(
data,
groups = NULL,
fill_area = TRUE,
cols = NULL,
chrs = paste0("chr", c(1:22, "X")),
genome_build = c("hg19", "hg38", "T2T", "mm10", "mm9", "ce11"),
cutoff = 2L,
resolution_factor = 1L,
force_y_limit = TRUE,
highlight_genes = NULL,
repel = FALSE,
nrow = NULL,
ncol = NULL,
return_plotlist = FALSE
)Arguments
- data
a
CopyNumberobject or a data.frame containing at least 'chromosome', 'start', 'end', 'segVal', 'sample' these columns.- groups
a named list or a column name for specifying groups.
- fill_area
default is
TRUE, fill area with colors.- cols
length-2 colors for AMP and DEL.
- chrs
chromosomes start with 'chr'.
- genome_build
genome build version, used when
datais adata.frame, should be 'hg19' or 'hg38'.- cutoff
copy number value cutoff for splitting data into AMP and DEL. The values equal to cutoff are discarded. Default is
2, you can also set a length-2 vector, e.g.c(2, 2).- resolution_factor
an integer to control the resolution. When it is
1(default), compute frequency in each cytoband. When it is2, use compute frequency in each half cytoband.- force_y_limit
default is
TRUE, force multiple plots- highlight_genes
gene list to highlight. have same y ranges. You can also set a length-2 numeric value.
- repel
if
TRUE(default isFALSE), repel highlight genes to avoid overlap.- nrow
number of rows in the plot grid when multiple samples are selected.
- ncol
number of columns in the plot grid when multiple samples are selected.
- return_plotlist
default is
FALSE, ifTRUE, return a plot list instead of a combined plot.
Value
a (list of) ggplot object.
Examples
load(system.file("extdata", "toy_copynumber.RData",
package = "sigminer", mustWork = TRUE
))
p1 <- show_cn_group_profile(cn)
p1
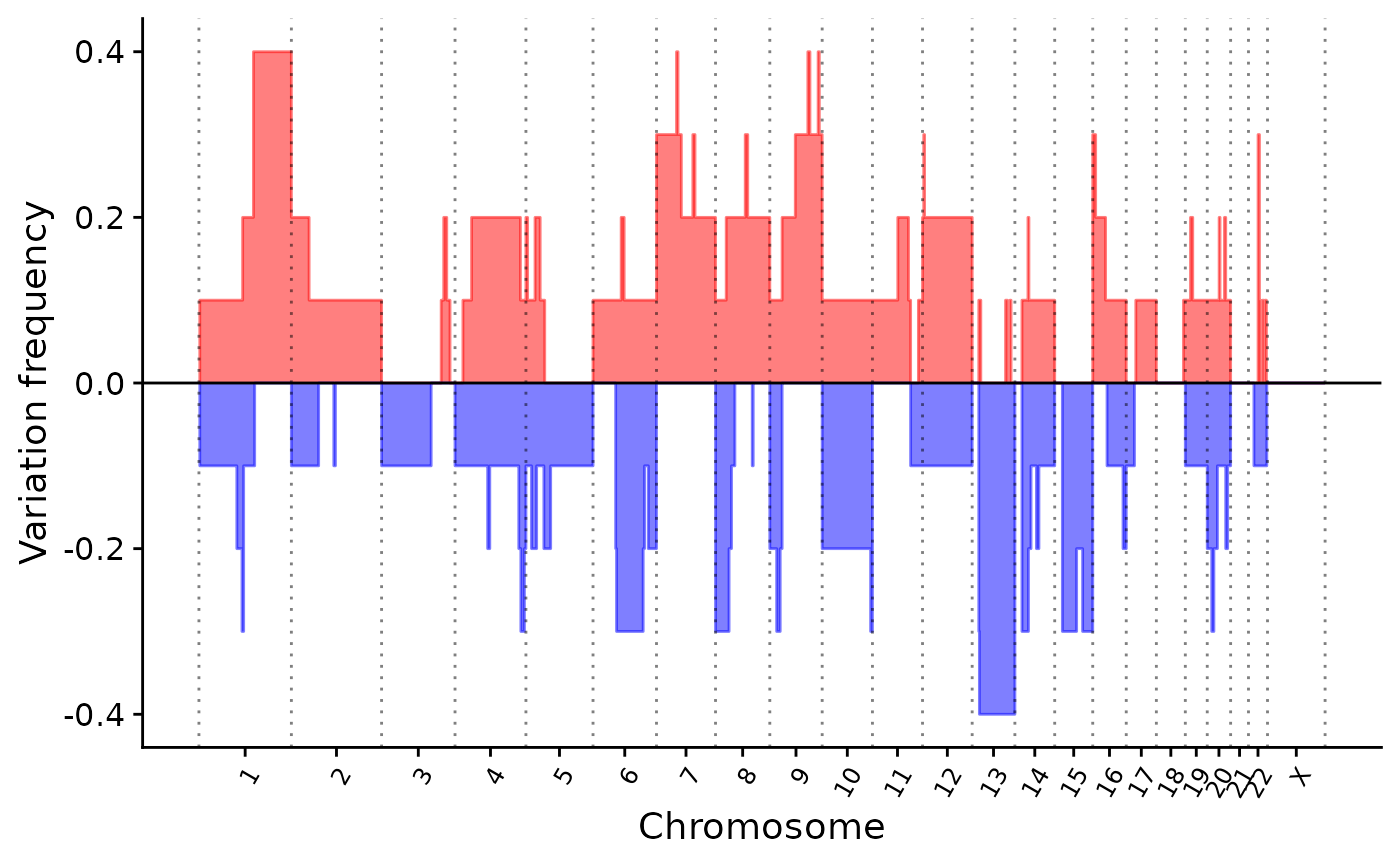 # \donttest{
ss <- unique(cn@data$sample)
p2 <- show_cn_group_profile(cn, groups = list(a = ss[1:5], b = ss[6:10]))
p2
# \donttest{
ss <- unique(cn@data$sample)
p2 <- show_cn_group_profile(cn, groups = list(a = ss[1:5], b = ss[6:10]))
p2
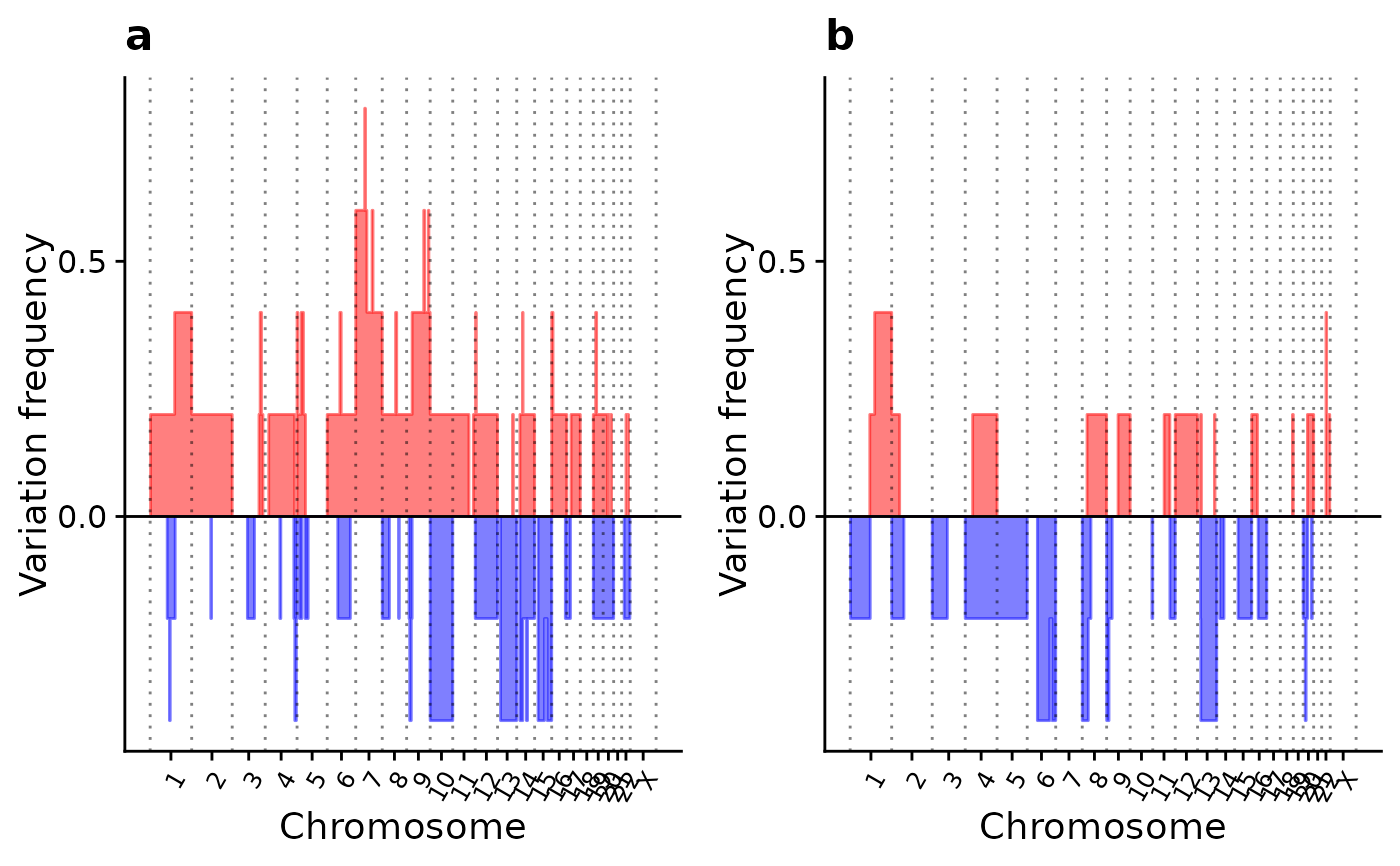 p3 <- show_cn_group_profile(cn,
groups = list(g1 = ss[1:5], g2 = ss[6:10]),
force_y_limit = c(-1, 1), nrow = 2
)
p3
p3 <- show_cn_group_profile(cn,
groups = list(g1 = ss[1:5], g2 = ss[6:10]),
force_y_limit = c(-1, 1), nrow = 2
)
p3
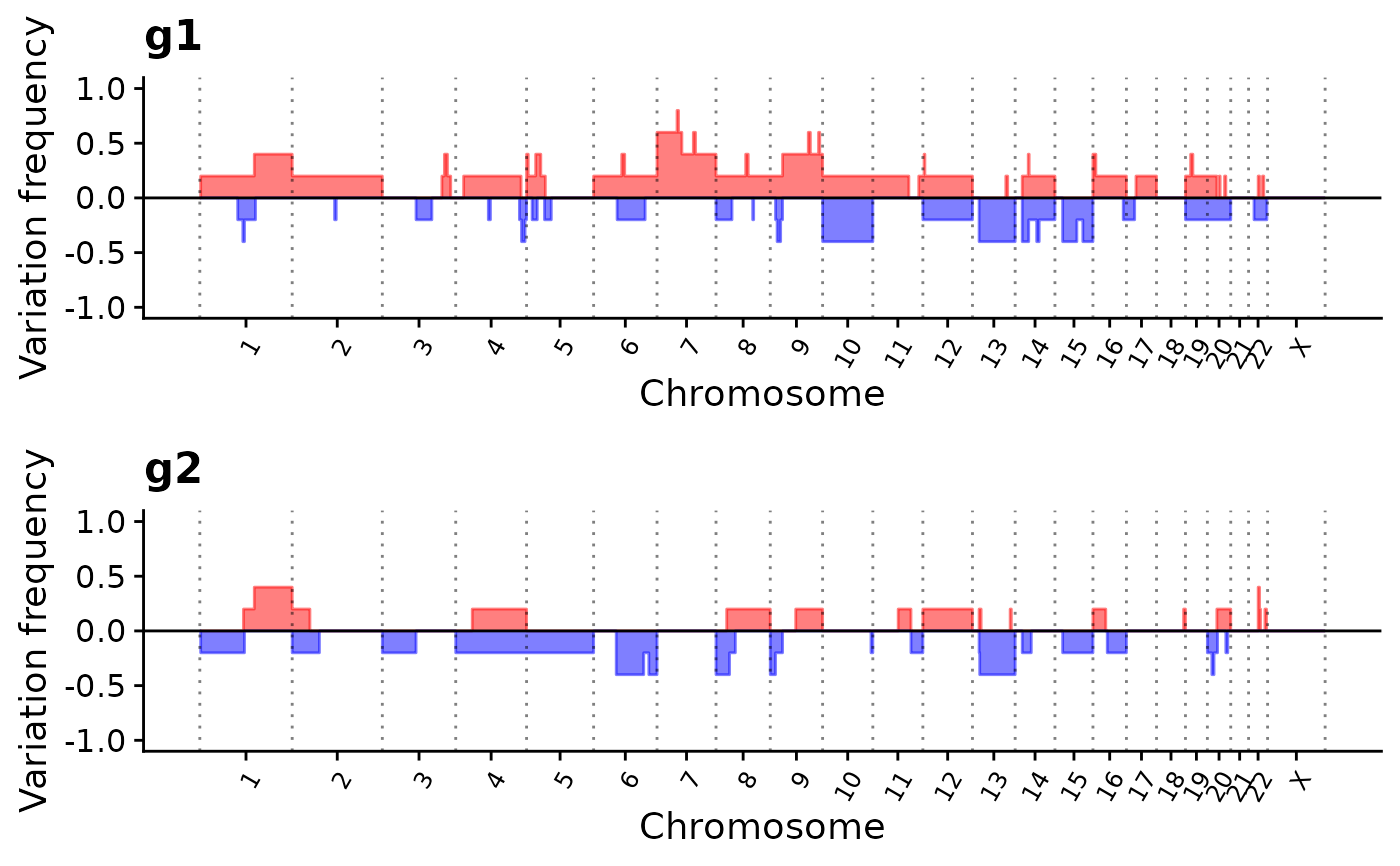 ## Set custom cutoff for custom data
data <- cn@data
data$segVal <- data$segVal - 2L
p4 <- show_cn_group_profile(data,
groups = list(g1 = ss[1:5], g2 = ss[6:10]),
force_y_limit = c(-1, 1), nrow = 2,
cutoff = c(0, 0)
)
p4
## Set custom cutoff for custom data
data <- cn@data
data$segVal <- data$segVal - 2L
p4 <- show_cn_group_profile(data,
groups = list(g1 = ss[1:5], g2 = ss[6:10]),
force_y_limit = c(-1, 1), nrow = 2,
cutoff = c(0, 0)
)
p4
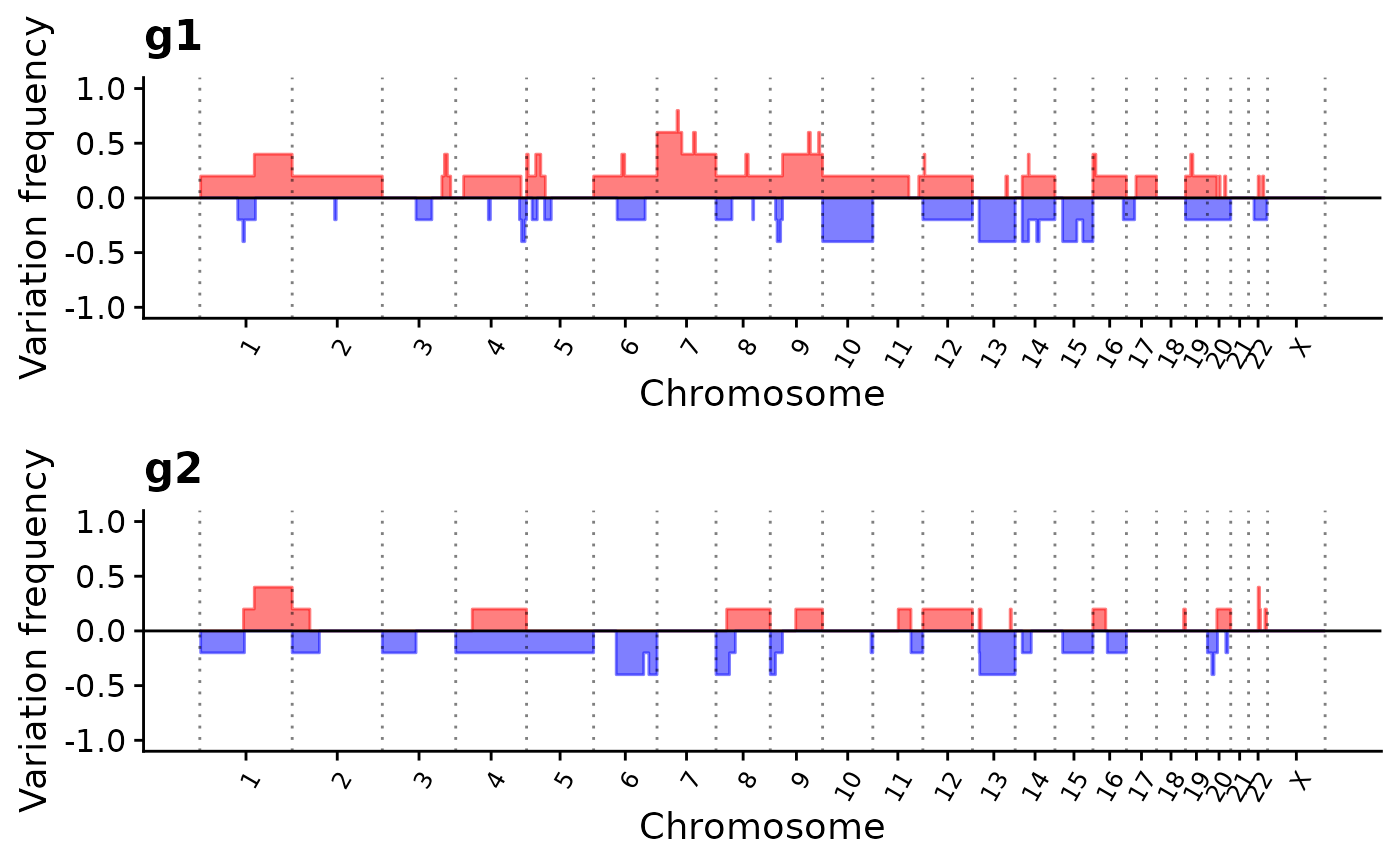 ## Add highlight gene
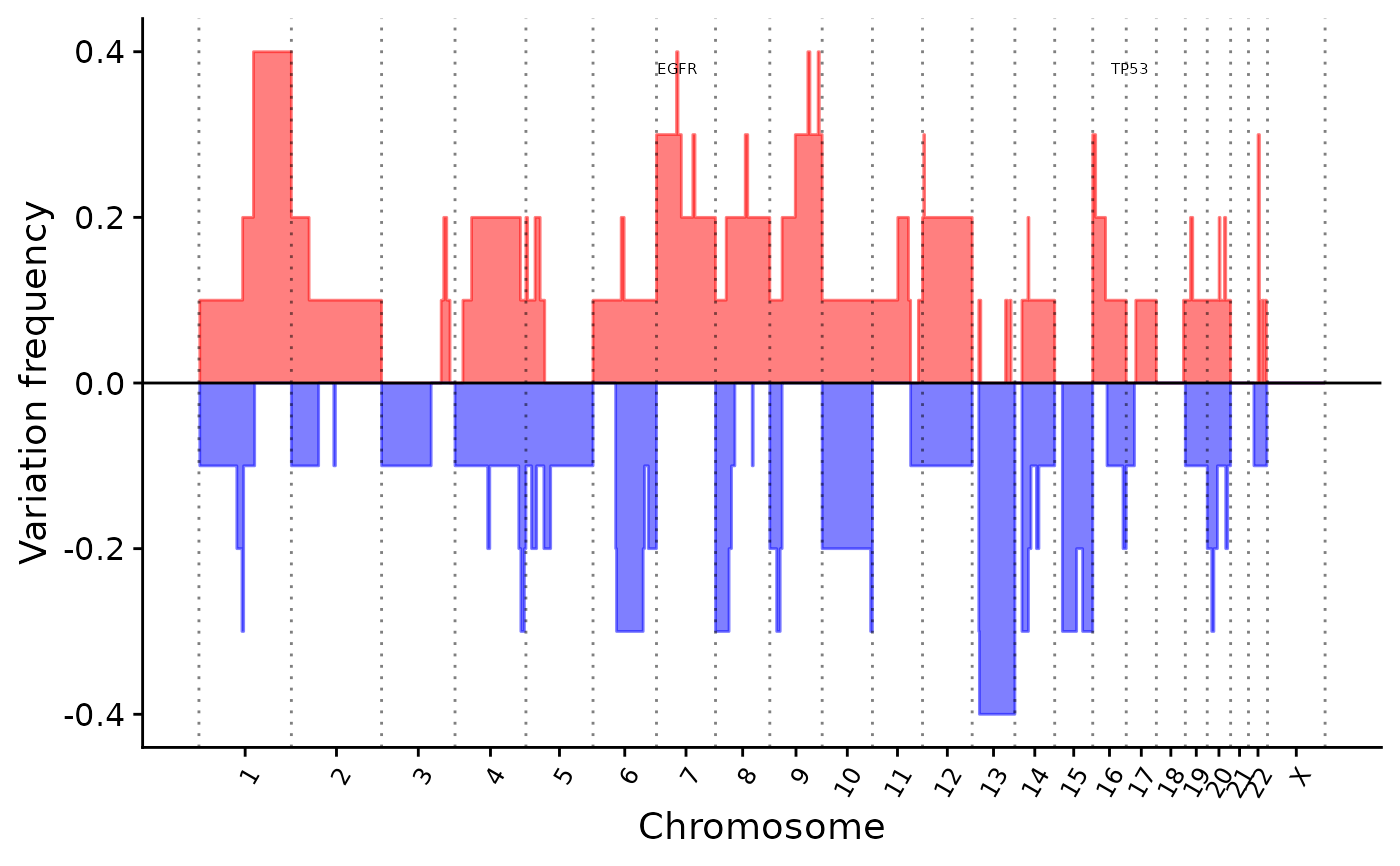
p5 <- show_cn_group_profile(cn, highlight_genes = c("TP53", "EGFR"))
p5
## Add highlight gene
p5 <- show_cn_group_profile(cn, highlight_genes = c("TP53", "EGFR"))
p5
 # }
# }